|
12/8/2023 0 Comments Cytoscape linux  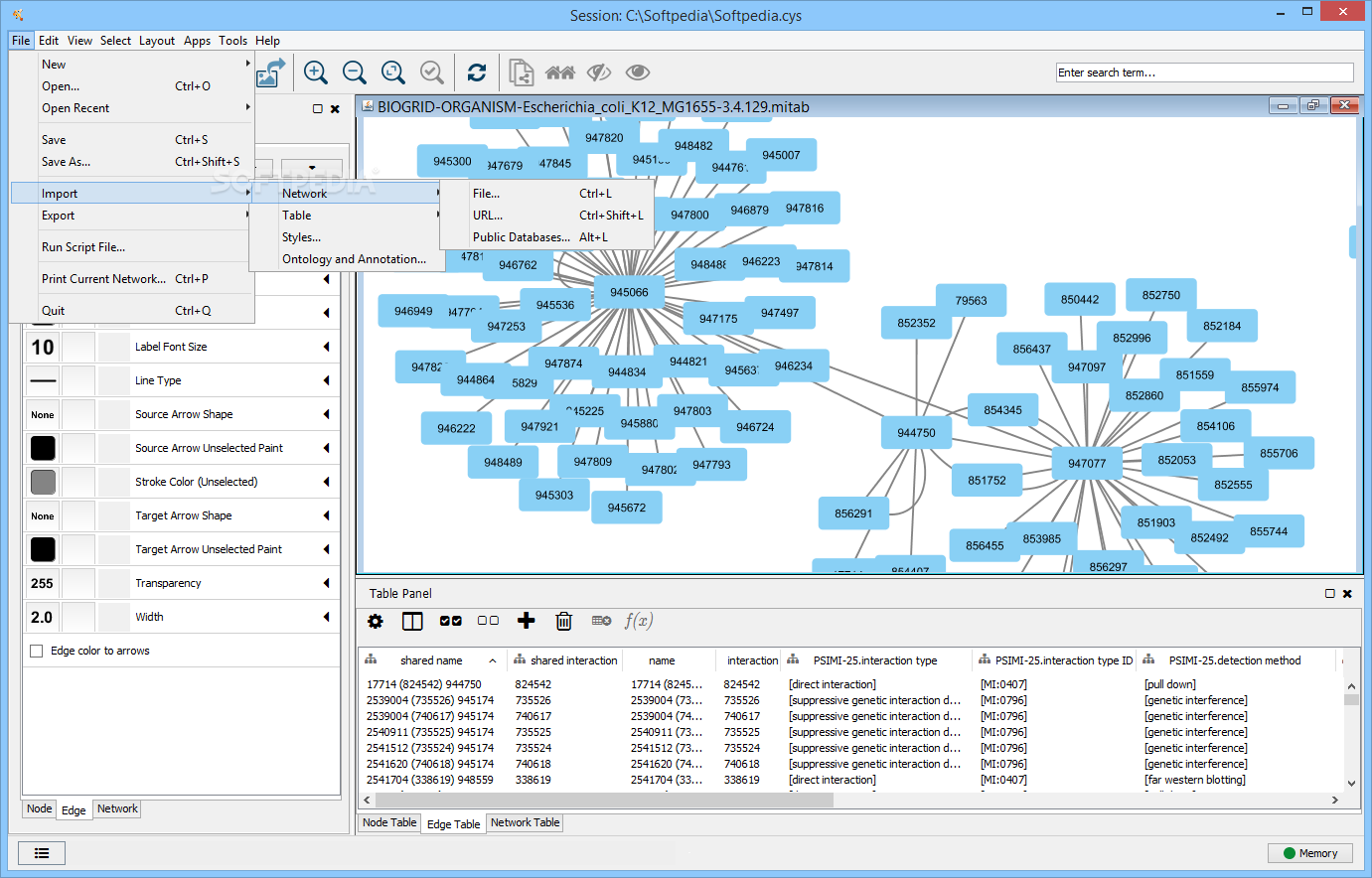
CABIN: collective analysis of biological interaction networks. Estimating and improving protein interaction error rates. How reliable are experimental protein–protein interaction data? J. Weighing our measures of gene expression. GoMiner: a resource for biological interpretation of genomic and proteomic data. BiNGO: a Cytoscape plugin to assess overrepresentation of gene ontology categories in biological networks. MAPPFinder: using gene ontology and GenMAPP to create a global gene-expression profile from microarray data. 
Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Functional genomics and proteomics: charting a multidimensional map of the yeast cell. Global studies of cell type-specific gene expression in plants. Global survey of organ and organelle protein expression in mouse: combined proteomic and transcriptomic profiling. This should solve the problem.Kislinger, T. If it is not, remove the existing BiNGO files and unzip the BiNGO zip file again, directly into the /plugins directory, not into /plugins/bingo or such. BiNGO assumes the following directory structure : there should be a file called BiNGO.jar and a folder called BiNGO (containing the annotation files) in the /plugins directory of Cytoscape (produced by unzipping the BiNGO zip file in the plugins folder). Apparently this exception is not properly handled by BiNGO (it will be fixed in a later version). What is going on ?Ī : The fact that BiNGO specifically stalls at 0% indicates that the program can't find the annotation and ontology files in the proper directory. 
Q : BiNGO stalls at 0% while parsing annotations. if you want to change the default organism to A. Can I do that ?Ī : Yes, you can modify most of BiNGO's default settings by modifying the bingo_gui.properties file in the BiNGO.jar archive. Q : I would like to change the default settings in the BiNGO Settings Panel. You can use these as custom annotation/ontology files. I would like to use more recent annotations.Ī : Download the most recent annotation and ontology files from the GO website. Q : The default annotations/ontologies in BiNGO are already several months old. However, these genes do have an annotation and they are taken into account. This means that some curator has actually looked at them and had to conclude that there is as yet no clear-cut functional classification. There is however a difference with genes that are annotated to categories such as Biological_Process unknown. The genes for which there is no annotation available are not taken into account in the over-representation analysis, and they're not counted as genes belonging to the test or reference set. Given the current status of GO annotation efforts, a lot of genes haven't actually been looked at by a curator and annotated in the GO hierarchy. Q : The number of genes in my test set according to the BiNGO output window does not match with the number of nodes I pasted into the text field or selected in the graph. Try running Cytoscape from cytoscape.bat (Windows) or cytoscape.sh (Linux, Mac), or see here to increase Cytoscape’s default memory settings. Q : BiNGO aborts without any error message.Ī : Cytoscape’s default memory settings (particularly in version 2.7) are probably not sufficient. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
